DNA Data Storage: Unlocking Massive Information Capacity
Insights | 11-08-2020 | By Liam Critchley

Key Things to Know:
- Incredible Storage Capacity: A single strand of DNA can store up to 215 million gigabytes of data, making it one of the highest density storage mediums available.
- DNA Replication: DNA replication in eukaryotic cells occurs in the nucleus, with the enzyme DNA helicase playing a crucial role by unwinding the DNA double helix.
- Future of Data Storage: Scientists are exploring the vast potential of DNA for data storage due to its high capacity, long-term stability, and energy efficiency.
- DNA vs. RNA: While RNA is essential for carrying genetic instructions, DNA's stability and storage capacity make it a superior choice for long-term data storage solutions.
Data is at everyone’s fingertips in our current high-tech modern-day world, and the amount of data available to us all is exponentially increasing by the day. Over the years, and throughout the data boom age of the last decade or so, companies have sought ways of storing more significant amounts of data (both in hardware and in the cloud) and have looked at ways of storing large amounts of data in smaller hardware architectures.
The question arises: how much data is in a strand of DNA? Each strand can store an impressive amount of information, equivalent to millions of gigabytes, showcasing the incredible potential of DNA as a data storage medium. However, as the world continues to create more and more data, it needs to be stored somewhere. If the production of data carries on at the same (or greater) rate than we are currently producing, the amount of data will soon outpace the available data storage capabilities, so scientists and engineers are still trying to discover new cutting-edge data storage methods.
Data Storage within DNA
Most people know about DNA (aka deoxyribonucleic acid) as the molecule that holds the information, i.e. the genetic codes, within our bodies. What some people don’t realise is that this equates to a seriously large amount of information being stored within a single biomolecule. This brings us to another intriguing question: how much data is stored in DNA across the entire human body? The cumulative storage capacity of human DNA far exceeds that of any artificial medium, pointing to exciting possibilities for future data storage technologies. Simply put, the information storage devices within our bodies are much more capable than we can currently create, so there has been a lot of focus in trying to harness the power and data storage capabilities of DNA for our own man-made data storage systems.
It’s not just the storage density of DNA that is appealing; it also has a long usable lifetime and has a very good energy efficiency compared to other storage mediums. How much information can be stored in DNA? Scientists estimate that DNA's storage capacity allows it to hold billions of terabytes of data, making it a formidable medium for long-term data preservation. To read the data stored within these devices, the DNA strands are read by a DNA sequencer and the information held within the strands is decoded back into an electronically compatible form.
As it stands, a lot of the databases used in DNA-based storage devices are static, but many believe that these devices could include some dynamic aspects, and this could push the capabilities beyond what is currently possible. These dynamic implementations could be the key to realising commercially viable DNA-based storage systems.
Current Challenges of Data Storage
While the possibilities of DNA data storage devices are enormous, there are a few areas that need to be optimised before they are suitable for widespread use in our information and data storage systems—but this is nothing new, as all technologies need to get through similar optimisation phases.
As it stands, there are some limitations to using DNA as storage material in man-made systems, and its use often involves some trade-offs and compromises to be made. These trade-offs tend to be around the dynamic aspects of the storage devices, such as the physical scalability, encoding density or the reusability of the devices, as creating a device with all three capabilities has been a sticking point in DNA storage device designs.
For example, if we take the most common DNA-based storage-based method—which is the polymerase chain reaction (PCR) storage method. Another important aspect of DNA storage is understanding how much memory DNA has in practical applications. Research continues to focus on optimizing these systems to harness the full memory potential of DNA without compromising access frequency—and this is a reason that DNA-based storage devices have been touted for cold storage devices that don’t require regular access. It also requires the double-stranded DNA molecules to be melted in data payload regions, which means that while it can be encoded efficiently, the storage density suffers as a result.
A key consideration in these developments is determining how much data can human DNA store. With the ability to encode vast amounts of data into the double helix, human DNA represents a near-limitless storage resource.
A New DNA-based Data Storage Device
A team from the USA have now proffered a solution that could help to solve the trade-off issues between the dynamic aspects within DNA storage devices. Inspired by microbiological and genetic research, the researchers created the device so that it would function in a similar way to how cells access information from their genome, i.e. so the man-made system is not too far removed from the natural process.
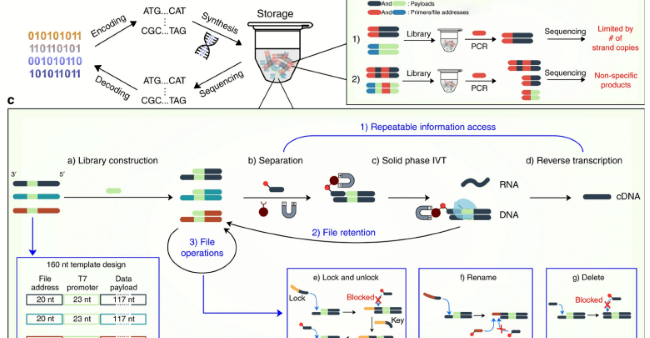
Credit: Nature Communications, Article number: 2981 (2020)
Understanding the role of enzymes like DNA helicase is crucial. But what type of biological molecule is DNA helicase? It's a protein that unwinds the DNA double helix, facilitating the processes of replication and transcription. Each of these strands were used to store information, and data storage systems could be constructed using many of these strands. Additionally, when we explore how much information is stored in DNA, we find that the complexity and density of genetic information make DNA a superior medium for both biological and digital data storage. Moreover, the overhang strand section allows each of the files to be separated from each other and allows in-storage operations to take place. These features have been made possible by the single strand having a T7 promoter (which is a sequence of nucleic acid bases) that enables the transcription of information into RNA at the same time, leaving the information stored in the double-stranded DNA.
This work is in its infancy and has only just proven that it is possible to create DNA-based information storage systems that are dynamic and without compromising on crucial functioning areas. Nevertheless, the work now sets a platform to build upon for building the next generation of versatile and high-density data storage devices. Moreover, the debate of RNA vs DNA often surfaces in the context of data storage. While RNA serves as a messenger in biological systems, DNA's superior stability and capacity make it the preferred medium for data storage innovations.

